from datasets import load_datasetMedSAM inference
MedSAM inference will be done here
dataset = load_dataset("nielsr/breast-cancer", split="train")idx = 10
image = dataset[idx]["image"]
label = dataset[idx]["label"]
msk = np.array(label)
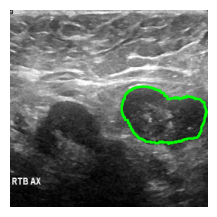
an_img = overlay_mask_border_on_image_frm_img(
image, msk,
)
show_(an_img)
#msk.shape, image.size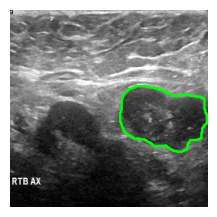
cntrs = find_contours_binary(msk.astype(np.uint8))[0]
x, y, w, h = frm_cntr_to_bbox(cntrs)def get_model():
model = SamModel.from_pretrained("wanglab/medsam-vit-base")
return modelget_bounding_box
get_bounding_box (ground_truth_map:numpy.ndarray)
Get bounding box from mask
get_prediction
get_prediction (model:transformers.models.sam.modeling_sam.SamModel, model_name:str, image:PIL.Image.Image, boxes:List[int], device:Optional[str]=None, threshold:float=0.5)
| Type | Default | Details | |
|---|---|---|---|
| model | SamModel | ||
| model_name | str | checkpoint in hugggingface | |
| image | Image | ||
| boxes | List | ||
| device | Optional | None | |
| threshold | float | 0.5 | |
| Returns | Tuple |
model = get_model()preds_ = get_prediction(
model=model,
model_name='wanglab/medsam-vit-base',
image=image,
boxes=[get_bounding_box(msk)],
device='cpu',
threshold=0.9
)an_img = overlay_mask_border_on_image_frm_img(image, preds_)
show_(an_img)